 常用生物信息学数据库和分析工具网址Word格式文档下载.doc
常用生物信息学数据库和分析工具网址Word格式文档下载.doc
- 文档编号:4642696
- 上传时间:2023-05-03
- 格式:DOC
- 页数:10
- 大小:250KB
常用生物信息学数据库和分析工具网址Word格式文档下载.doc
《常用生物信息学数据库和分析工具网址Word格式文档下载.doc》由会员分享,可在线阅读,更多相关《常用生物信息学数据库和分析工具网址Word格式文档下载.doc(10页珍藏版)》请在冰点文库上搜索。
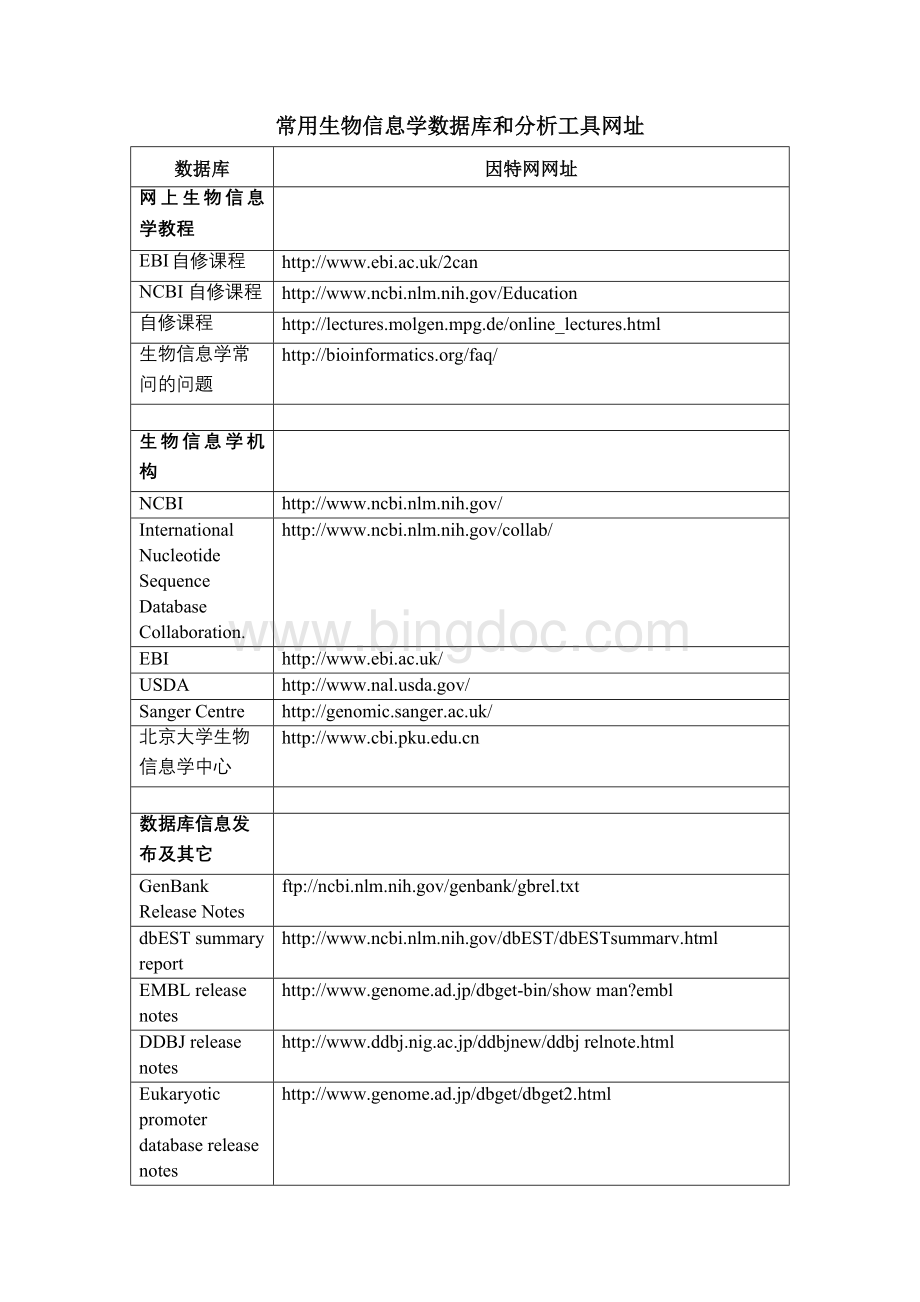
Eukaryoticpromoterdatabasereleasenotes
//www.genome.ad.jp/dbget/dbget2.html
SwissProtreleasenotes
swissprot
PIRreleasenotes
pir
PRFreleasenotes
prf
PDBSTRreleasenotes
pdbstr
Prositereleasenotes
prosite
PDBreleasenotes
pdb
KEGGreleasenotes
pathway
核苷酸数据库
GenBank
dbEST
//www.ncbi.nlm.nih.gov/dbEST/index.html
dbSTS
//www.ncbi.nlm.nih.gov/dbSTS/index.html
dbGSS
//www.ncbi.nlm.nih.gov/dbGSS/index.html
Genome(NCBI)
//www.ncbi.nlm.nih.gov/entrez/query.fcgi?
db=Genome
dbSNP
//www.ncbi.nlm.nih.gov/SNP/
HTGS
//www.ncbi.nlm.nih.gov/HTGS/
UniGene
//www.ncbi.nlm.nih.gov/UniGene/
EMBL核苷酸数据库
//www.ebi.ac.uk/embl
Genome(EBI)
//www.ebi.ac.uk/genomes/
向EMBL数据库提交序列
//www.ebi.ac.uk/embl/Submission/webin.html
DDBJ
//www.ddbj.nig.ac.jp/
PlantRgenedatabase
//www.ncgr.org/rgenes
启动子数据库
Eukaryoticpromoterdatabase
//www.epd.isb-sib.ch
转录因子数据库
FRANSFAC
//transfac.gbf.de
ooTFD
//www.ifti.org
基因注释数据库
RAP-DB
//rapdb.lab.nig.ac.jp
基因分类数据库
GeneOntology(GO)
//www.geneontology.org
蛋白质数据库
SWISS-PROT或TrEMBL
//www.ebi.ac.uk/swissprot/
//www.expasy.ch/sprot/
PIR
//pir.georgetown.edu
PRF
//www.kinasenet.org/pkr/Welcome.do
//www.prf.or.jp/
PDBSTR
//www.genome.ad.jp
Prosite
//www.expasy.org/prosite
结构数据库
PDB
//www.rcsb.org/pdb
//www.pdb.org
NDB
//ndbserver.rutgers.edu/NDB/ndb.html
//ndbserver.rutgers.edu/
DNA-BindingProteinDatabase
//ndbserver.rutgers.edu/NDB/structure-finder/dnabind/index.html
NMRNucleicAcidsDatabase
//ndbserver.rutgers.edu/NDB/structure-finder/nmr/index.html
ProteinPlusDatabase
//ndbserver.rutgers.edu/NDB/structure-finder/protein/index.html
Swiss3Dimage
//www.expasy.ch/sw3d/
SCOP
//scop.mrc-lmb.cam.ac.uk/scop/
CATH
//www.biochem.ucl.ac.uk/bsm/cath/
酶、代谢和调控路径数据库
KEGG
//www.genome.ad.jp/kegg/
EnzymeNomenclatureDatabase
//expasy.hcuge.ch/sprot/enzyme.html
ProteinKinaseResource(PKR)
//www.sdsc.edu/kinases/
LIGAND
//www.genome.ad.jp/dbget/ligand.html
WIT
//www.cme.msu.edu/WIT/
EcoCyc
//ecocyc.PangeaS
UM-BBD
//www.labmed.umn.edu/umbbd/
多种代谢路径数据库
//www.unl.edu/stc-95/ResTools/biotools/biotools8.html
基因调控路径数据库(TRANSPATH)
基因组数据库
禾本科比较基因组
//www.gramene.org
GrainGene
//www.graingenes.org
BotanicalData
//www.calflora.org/calflora/batanical.html
日本水稻基因组(RGP)
//rgp.dna.affrc.go.jp
水稻物理图谱
//www.genome.clemson.edu/projects/rice/fpc
//www.genome.arizona.edu
华大水稻基因组框架图
欧洲水稻测序(第12染色体)
s.fr
OryGenesDB(水稻插入突变体)
//orygenesdb.cirad.fr
Maizegenome
//www.agron.missouri.edu
Barleygenome
//www.css.orst.edu/Research/barley/nabgmp.htm
Foragegrassesgenomes
//forages.orst.edu/
//www.forages.css.orst.edu/Topics/Species/Grasses/
Triticumgenomes
//wheat.pw.usda.gov/index.shtml
Arabidopsisgenome
//www.arabidopsis.org
SoyBase
//soybase.agron.iastate.edu
Alfalfagenome
//www.alfalfa.ksu.edu
Cottongenome
//cottongenomecenter.ucdavis.edu
Glycinemaxgenome
//www.zmdb.iastate.edu/PlantGDB/glycine_max.html
//www.zmdb.iastate.edu/PlantGDB
C.elegansgenome
//www.acedb.org
藻类(Chlamydomonas)基因组
//www.biology.duke.edu/chlamy_genome
粘菌(Dictyostelium)基因组
//dictygenome.bcm.tmc.edu
Animalgenomes(ArkDB)
//www.thearkdb.org
FlyBase
//flybase.bio.indiana.edu/.bin/fbidq.html?
FBgn0003075
MouseGenomeInformatics
//www.informatics.jax.org/bin/query_accession?
id=MGI:
97555
SaccharomycesGenomeDatabase
//genome-www.stanford.edu/cgi-bin/dbrun/SacchDB?
find+Locus+%22PGK1%22
多种基因组数据库
//www.hgmp.mrc.ac.uk/GenomeWeb
RiceMutantDatabase
文献数据库
PubMed
//www.ncbi.nlm.nih.gov/PubMed/
OMIM
//www.ncbi.nlm.nih.gov/Omim/
Agricola
//www.nal.usda.gov/ag98/
RiceGeneticsNewsletter
//www.gramene.org/newsletters/rice_genetics
ProceedingsoftheNationalAcademyofSciencesUSA(PNAS)
//intl.pnas.org
关键词为基础的数据库检索
Entrez
//www.ncbi.nlm.nih.gov/Entrez/
EntrezNucleotideSequenceSearch
//www.ncbi.nlm.nih.gov/Entrez/nucleotide.html
EntrezProteinSequenceSearch
//www.ncbi.nlm.nih.gov/Entrez/protein.html
BatchEntrez
//www.ncbi.nlm.nih.gov/Entrez/batch.html
SequenceRetrievalSystem,India
//bioinfo.ernet.in:
80/srs5/
SequenceRetrievalSystem,Singapore
//www.bic.nus.edu.sg:
SequenceRetrievalSystem,US
//iubio.bio.indiana.edu:
80/srs/srsc
SequenceRetrievalSystem,UK
//srs.ebi.ac.uk/
GetEntryNucleotide&
ProteinSequenceSearch
//ftp2.ddbj.nig.ac.jp:
8000/getstart-e.html
DatabaseSearchwithKeyWords
8080/dbsearch-e-new.html
DBGET/LinkDB
//www.genome.ad.jp/dbget/
序列为基础的数据库检索
BLAST
//www.ncbi.nlm.nih.gov/BLAST/
FASTA
//www.ebi.ac.uk/fasta33/index.html
BLITZ
//www2.ebi.ac.uk/bic_sw/
SSearch
rs.fr/bin/ssearch-guess.cgi
ElectronicPCR
//www.ncbi.nlm.nih.gov/STS/
Proteomeanalysis
//www.ebi.ac.uk/proteome/
Globalalignment
//genome.cs.mtu.edu
多序列分析
Clustalmultiplesequencealignment
//dot.imgen.bcm.tmc.edu
BCM
//dot.imgen.bcm.tmc.edu:
9331/multi-align/multi-align.html
EBIClustalW
//www.ebi.ac.uk/clustaw/index.html
修饰对序列对位排列结果的格式(Boxshade)
//www.ch.embnet.org/software/BOX_form.html
系谱分析
PAUP
//onyx.si.edu/PAUP/
EBIClustalWanalysis
//www.ebi.ac.uk
GCGpackage
PHYLIP
//evolution.genetics.washington.edu/phylip.html
MEGA/METREE
//www.bio.psu.edu/imeg
Hennig86
//www.vims.edu/~mes/hennig/software.html
GAMBIT
//www.lifesci.ucla.edu/mcdbio/Faculty/Lake/Research/Programs/
MacClade
//phylogeny.arizona.edu/macclade/macclade.html
Phylogeneticanalysis
//www.unl.edu/stc-95/ResTools/biotools/biotools2.html
ClustalX
//ftp-igbmc.u-strasbg.fr/pub/ClustalX
MEGA
TreeView
//taxonomy.zoology.gla.ac.uk/rod/treeview.html
基因结构预测分析
GENSCAN
//genes.mit.edu/GENSCAN.html
//bioweb.pasteur.fr/seqanal/interfaces/genscan-simple.html
//bioweb.pasteur.fr
GeneFinder
//www.bioscience.org/urllists/genefind.htm
GeneFinding
GeneFeatureSearches
Grail
//compbio.ornl.gov/Grail-1.3
GrailEXP
//compbio.ornl.gov/grailexp
GeneMark
//opal.biology.gatech.edu/GeneMark/eukhmm.cgi
//genemark.biology.gatech.edu/GeneMark/hmmchoice.html
Veil
//www.cs.jhu.edu/labs/compbio/veil.html
AAT
//genome.cs.mtu.edu/aat.html
GENEID
//www.imim.es/GeneIdentification/Geneid/geneid_input.html
Genlang
//cbil.humgen.upenn.edu/~sdong/genlang_home.html
GeneParser
//beagle.colorado.edu/~eesnyder/GeneParser.html
Glimmer
//www.cs.jhu.edu/labs/compbio/glimmer.html
MZEF
//www.cshl.org/genefinder
Procrustes
//www-hto.usc.edu/software/procrustes/
TandemRepeatsFinder
//C3.biomath.mssm.edu/trf.html
Repeats
//bioweb.pasteur.fr/seqanal/interfaces/repeats.html
基因分类
GOAnnotator
//udgenome.ags.udel.edu/gofigure
蛋白质结构预测分析
Expasy
//www.expasy.ch/
CBS
//www.cbs.dtu.dk
Predictingproteinsecondarystructure
9331/pssprediction/pssp.html
Predictingprotein3DStructures
//dove.embl-heidelberg.de/3D/
Predictingproteinstructures
9331/seq-search/struc-predict.html
其它分析工具和软件
BioEdit
//www.mbio.ncsu.edu/BioEdit/bioedit.html
Primer3(PCR引物设计)
//frodo.wi.mit.edu/cgi-bin/primer3/primer3_www.cgi
PutativeDNASequencingErrorsCheck
//www.bork.embl-heidelberg.de/Frame/
MatInspector
//www.gsf.de/cgi-bin/matsearch.pl
FastM
//www.gsf.de/cgi-bin/fastm.pl
WebSignalScan
//www.dna.affrc.go.jp/htdocs/sigscan/signal.html
BCMSearchLauncher
9331/seq-util/seq-util.html
Webcutter
TranslateDNAtoprotein
//www.expasy.ch/tools/dna.html
ABIM
//www-biol.univ-mrs.fr/english/logligne.html
sequencemotifs:
Pfam
//www.sanger.ac.uk/Pfam/
//pfam.wustl.edu/
- 配套讲稿:
如PPT文件的首页显示word图标,表示该PPT已包含配套word讲稿。双击word图标可打开word文档。
- 特殊限制:
部分文档作品中含有的国旗、国徽等图片,仅作为作品整体效果示例展示,禁止商用。设计者仅对作品中独创性部分享有著作权。
- 关 键 词:
- 常用 生物 信息学 数据库 分析 工具 网址
 冰点文库所有资源均是用户自行上传分享,仅供网友学习交流,未经上传用户书面授权,请勿作他用。
冰点文库所有资源均是用户自行上传分享,仅供网友学习交流,未经上传用户书面授权,请勿作他用。


 二年级下册数学专项练习-应用题1.docx
二年级下册数学专项练习-应用题1.docx
 红色精美二十届三中全会提出的新概念新观点新论断.pptx
红色精美二十届三中全会提出的新概念新观点新论断.pptx
